.png)
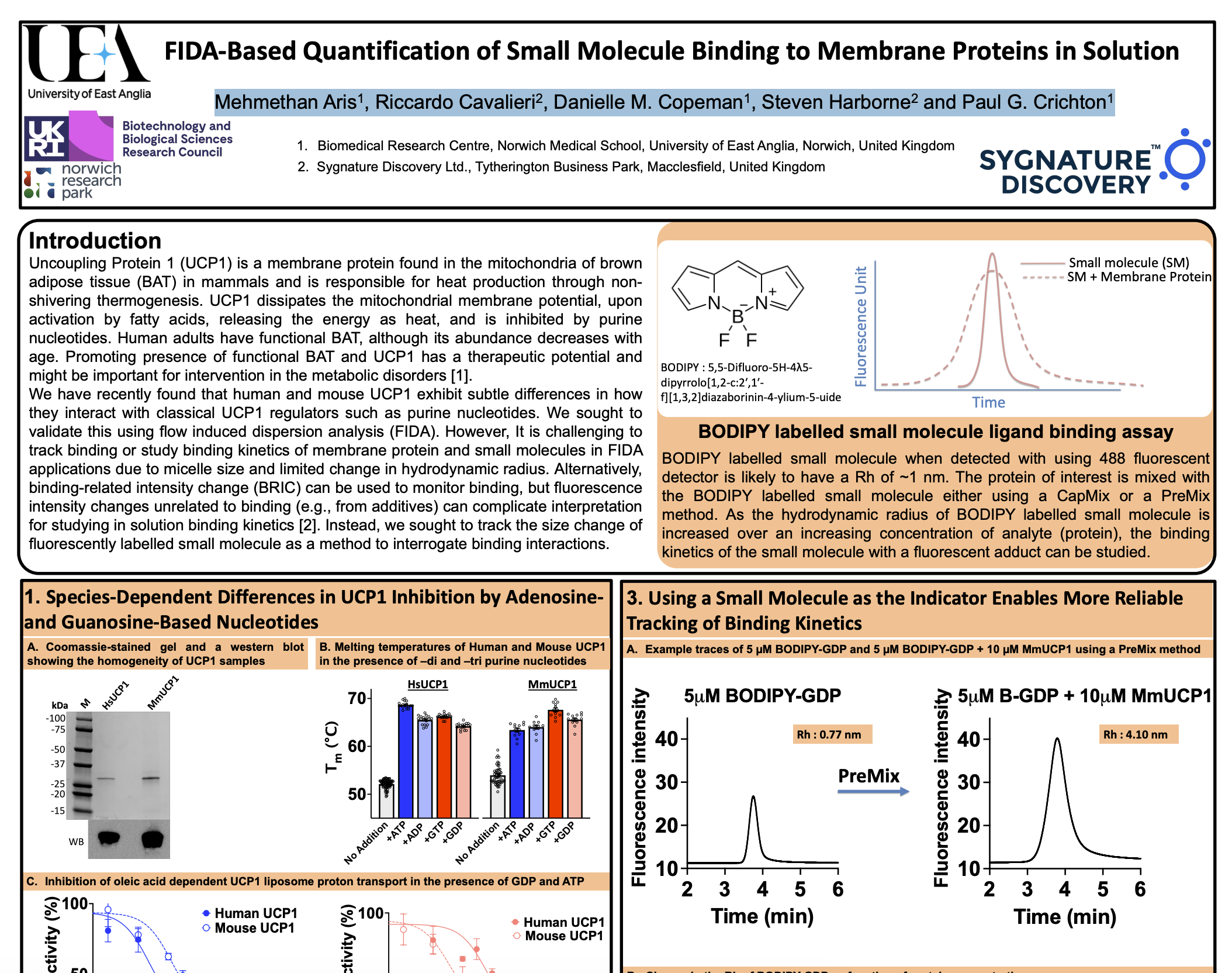
What does FIDA bring to Membrane Protein research?
01
Save sample material
- Down to 40 nl of sample
- No purification needed – high tolerance toward impurities
- No restriction on buffer composition
- Delivers data needed to optimised membrane protein expression conditions and increase yield
- Possible sample recovery
02
Tagged or label-free
- Tag flexibility: GFPs, HIS, Alpha tag, FLAG and similar
- Possibility of label-free characterisation of purified membrane protein
03
8 quality parameters in 1 run
- Assess structural changes
- Validate structures against reference models using built-in PDB Correlator (integration with Protein Data Bank)
- Know the absolutesize (Rh) measurement
- Determine your sample’s Polydispersity Index (PDI)
- Check for oligomeric state
- Quantify aggregation behaviour of your sample
- Detect ligand-binding and quantify Kd
04
Short assay optimisation and validation
- Save time with dedicated Assay Design tool
- First principle methods - allow you to avoid assumptions
- Built-in QC accelerates your optimisation process
- Absolute readouts and structural correlation facilitate efficient and reliable data validation
05
Temperature controlled – retain structure and function
- Keep your sample stored at 5°C while operating in walk-away mode
- Independent temperature control in sample and capillary compartment
Efficient analysis and rapid decision-making
With the ability to utilize sample volumes as low as 4μL (typically 10μL) for the analyte and 40 nL for the indicator, as well as the flexibility to operate directly in unpurified sample matrices, FIDA users can obtain a greater number of data points in a significantly reduced timeframe.
The users place unpurified samples in the autosampler’s 96 well plates or 2x50 vials, and after a few hours they are able to make an informed choice on which membrane protein to purify for further analysis.
Fast and easy – the FIDA way.

Down to
4μL
Analyte
40nL
Indicator Use
What is your most frequent challenge in the lab?


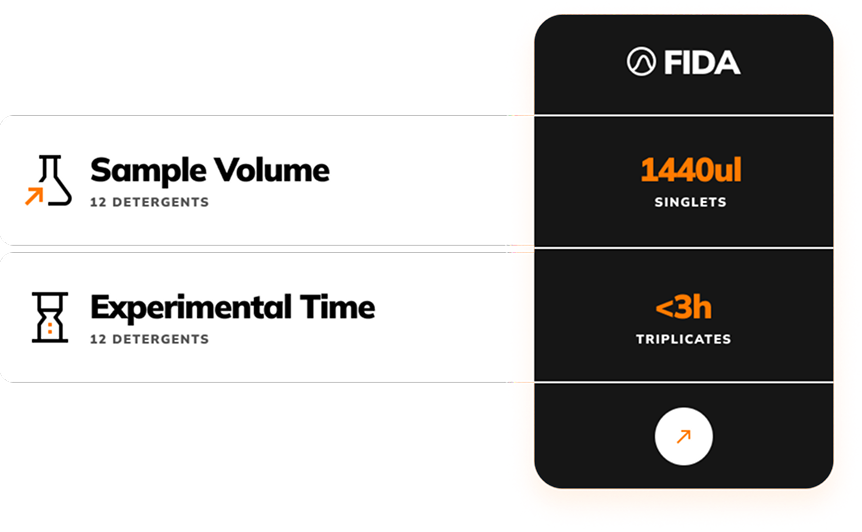
Detergent screening for solubilisation – 85% faster
Membrane Proteins Literature
Benefits

Keeps the cell lysate at 5°C while operating in walk-away mode

Quantitative detection of activation and binding in complex membrane constructs

Lower sample consumption

Speed to data

In-depth quality control

Informed troubleshooting
Features

Triple temperature control

Autosampler

Quantification of aggregates

Quantification of solubilised membrane protein

Polydispersity index

Oligomerisation
Measures

Accurate determination of dilute phase concentration

Relative droplet size distributions

Kinetics of droplet formation and maturation into amyloid fibrils

Determination of binding affinity between the polypeptide undergoing LLPS and LLPS-modulating compounds
Targeting common Membrane Protein Research Challenges

Challenge


Expression and purification
FIDA users can determine the in-solution concentration of membrane proteins. This allows them to optimise protein crystallisation conditions and to monitor protein expression and purification processes.
With just 0.12 uL per detergent (for triplicates) in detergent screening and down to 40nl of unpurified sample needed for characterisation, FIDA uses 10 to 100 less sample volume compared to other available technologies.
With just 0.12 uL per detergent (for triplicates) in detergent screening and down to 40nl of unpurified sample needed for characterisation, FIDA uses 10 to 100 less sample volume compared to other available technologies.
Stability and conformational heterogeneity.
Triple temperature control and possibility to work in unpurified matrices allows FIDA users to increase their level of control over membrane protein stability.
Functional characterisation
Working in crude matrices allows FIDA users to increase the applicability of their results. Additionally, the polydispersity index ensures that sample heterogeneity is considered in functional characterisation.
Structural characterisation
The size measurement delivered by FIDA allows our users to monitor changes as a result of interactions with other proteins or ligands. This information is then used to analyse the structure and function of membrane proteins of interest.
FIDA is currently used in cryo-EM sample preparation processes, as well as in nanobody clone selection. Our users can apply a single technology to characterise their membrane proteins, screen detergents, produce nanobodies to immobilise their membrane proteins and run cryo-EM sample preparation.
FIDA is currently used in cryo-EM sample preparation processes, as well as in nanobody clone selection. Our users can apply a single technology to characterise their membrane proteins, screen detergents, produce nanobodies to immobilise their membrane proteins and run cryo-EM sample preparation.
Effective drug development
FIDA measures the change in the size of the membrane protein complex when a ligand is bound, making it an optimal tool for evaluating protein-ligand interactions. This information allows FIDA users to identify potential ligands for drug discovery.



Become a user
Your laboratory instruments should serve you, not the other way around. We’re happy to help you.